The structure view window can be opened by clicking 'Viewer' in the 'View' menu of the main menu bar or the 'Viewer' shortcut of the tool bar. It can also be directly accessed by clicking on the molecule name of interest in the window 'Results' or the point of interest in the graphs of PCA and Correlation analysis.
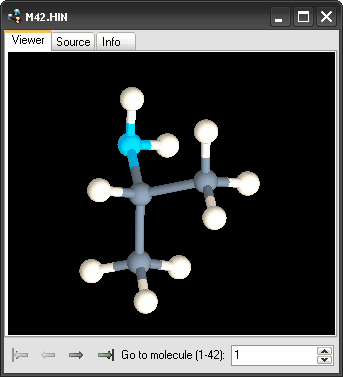
This window provides a graphical representation of the selected molecule. The name of the displayed molecule is reported as the window heading. The user can switch from one molecule to another by clicking the arrows at the bottom of the window or editing the molecule ID in the text box at the right.
The structure view window is comprised of three tabs. In the tab 'Viewer', the molecular structure of the selected molecule is shown. In the tab 'Source', the content of the molecular structure file of the selected molecule (or the part of it encoding the selected molecule) is shown. In the tab 'Info', a summary of the selected molecule is provided, it being comprised of the following:
| ▪ | Molecule Name (Name) |
| ▪ | Brutto Formula (Formula) |
| ▪ | Total number of atoms (Atom count) |
| ▪ | Total number of hydrogens (Hydrogen count) |
| ▪ | Total number of bonds (Bond count) |
| ▪ | Cyclomatic number (i.e., number of rings) |
| ▪ | Molecular mass (Mass) |
| ▪ | Filename (i.e., name and path of the input file encoding the selected molecule) |
| ▪ | Index of the molecule in input file (Index in file) |
| ▪ | Additional information, if present in the input file. |
It is possible to open a pop-menu with a right-click on the structure viewer. In this pop-up menu, different settings can be defined. One of this settings allows to change the molecule structure representation between original and Dragon bond orders. Note that the structure viewer in the load molecules window shows by default the original bonds defined in the molecule files.
Note that the tab Viewer is enabled only for molecular structure files where atom coordinates are defined. If atomic coordinates are missing as in SMILES notations, then only the Source and Info tabs will be enabled.